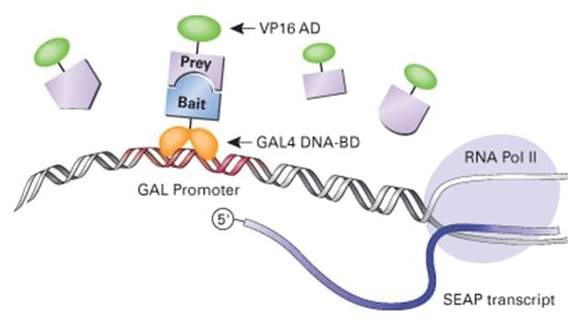
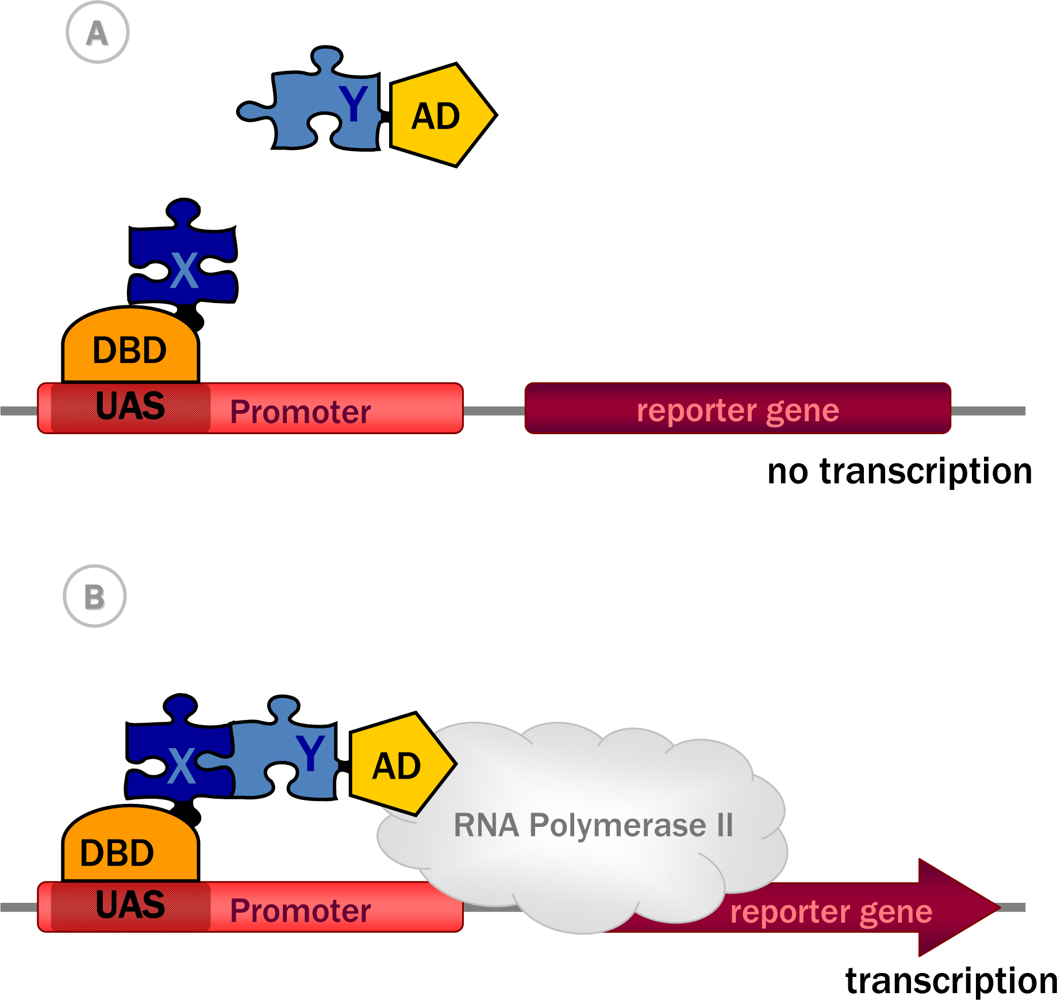
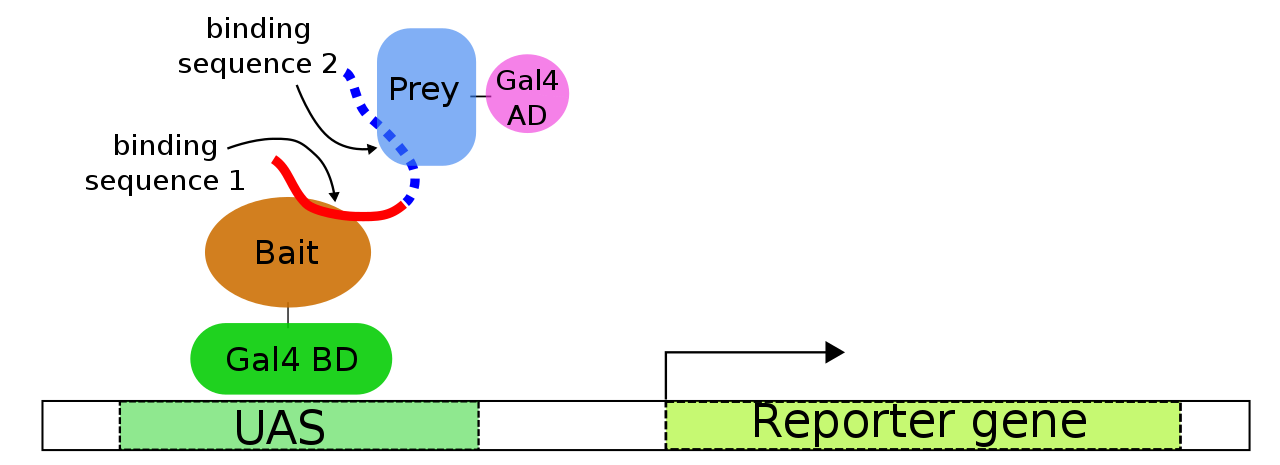
Diversity in Genetic In Vivo Methods for Protein-Protein Interaction Studies: from the Yeast Two-Hybrid System to the Mammalian Split-Luciferase System | Microbiology and Molecular Biology Reviews

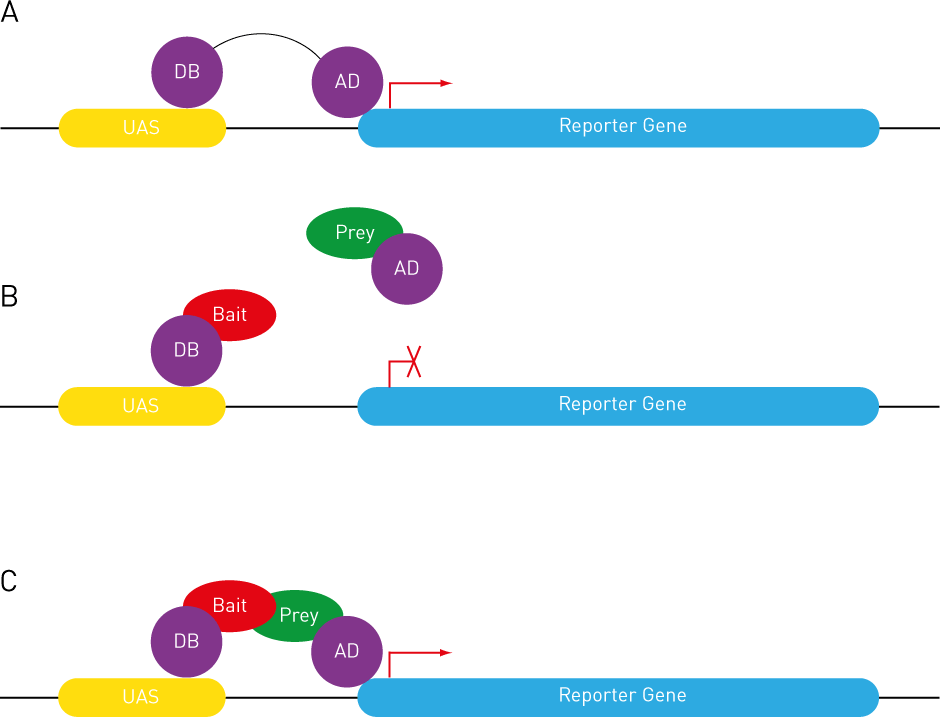
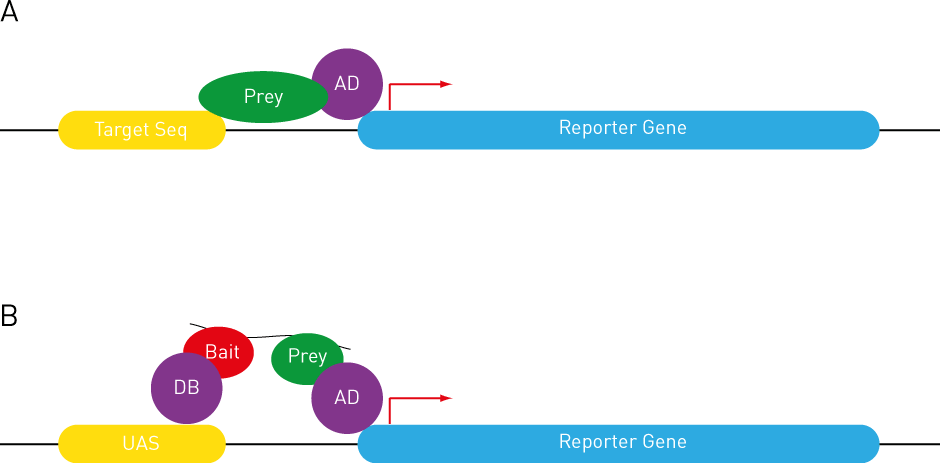
Reverse two-hybrid systems. (A) Repression of toxic genes (228, 373,... | Download Scientific Diagram
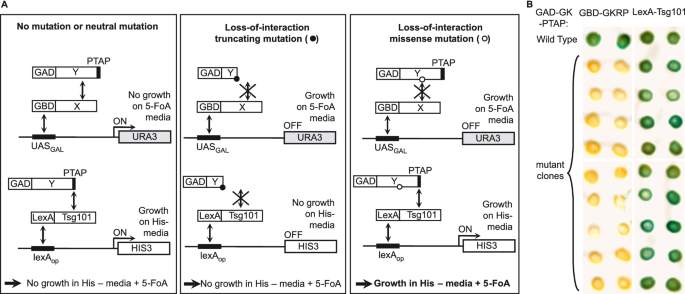
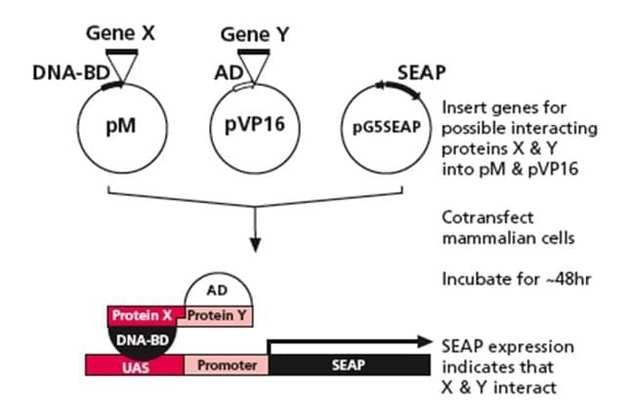
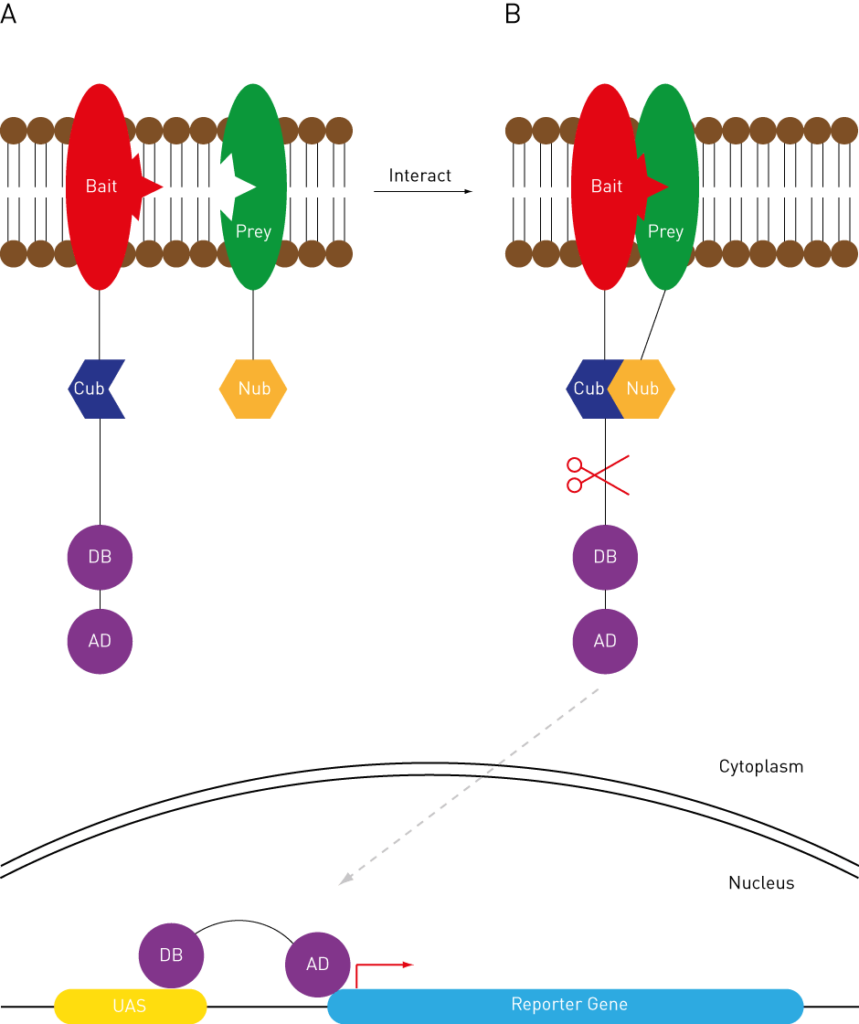
A novel reverse two-hybrid method for the identification of missense mutations that disrupt protein–protein binding | Scientific Reports
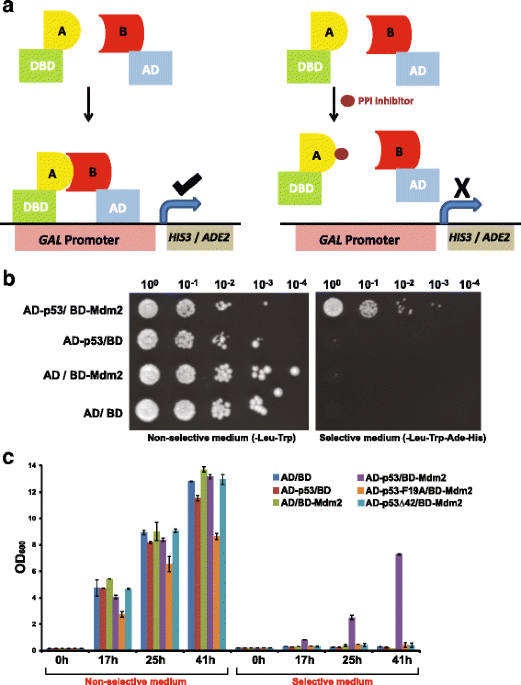
A yeast two-hybrid system for the screening and characterization of small-molecule inhibitors of protein–protein interactions identifies a novel putative Mdm2-binding site in p53 | BMC Biology | Full Text

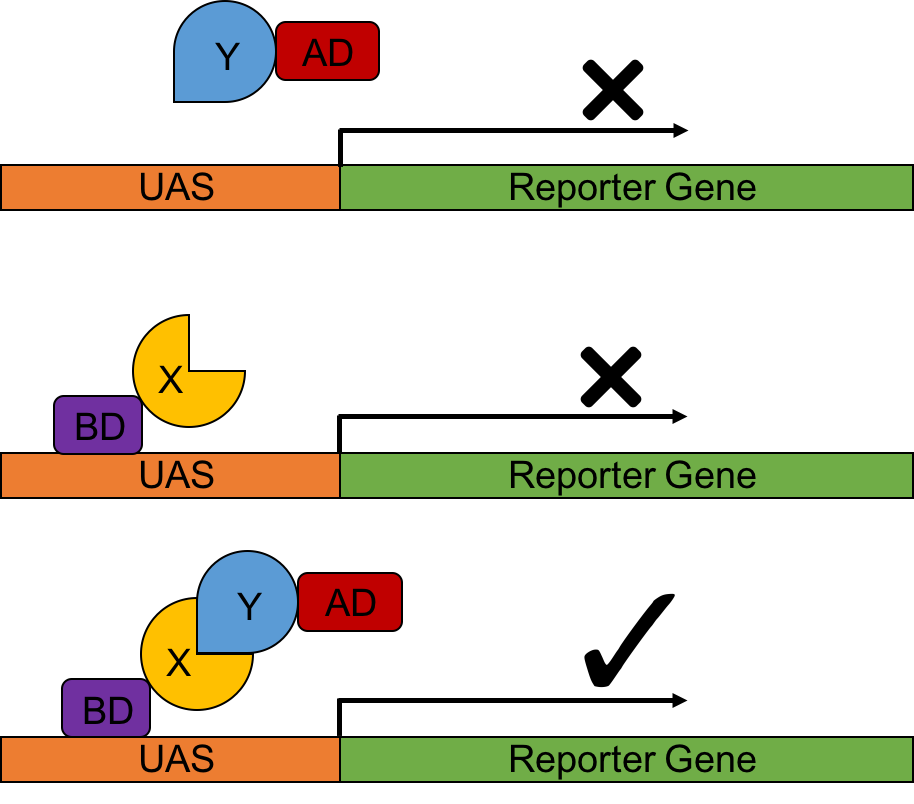
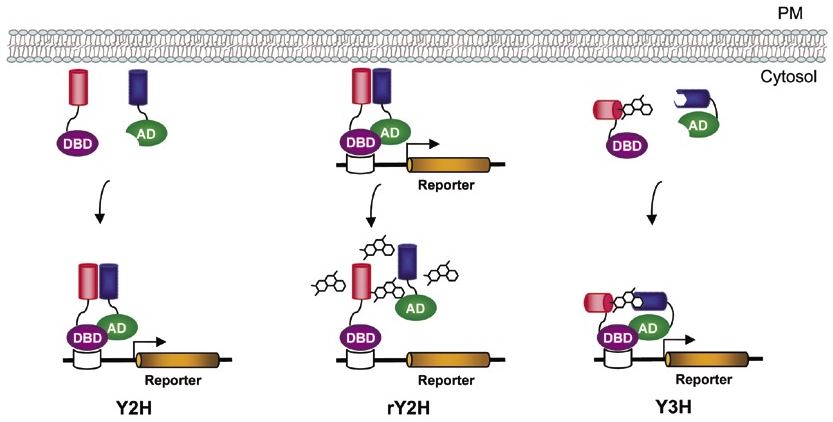
Figure 2 from A Comparison of Two-Hybrid Approaches for Detecting Protein-Protein Interactions. | Semantic Scholar

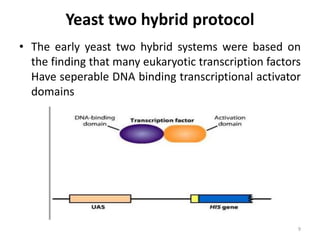
Yeast two-hybrid methods and their applications in drug discovery: Trends in Pharmacological Sciences